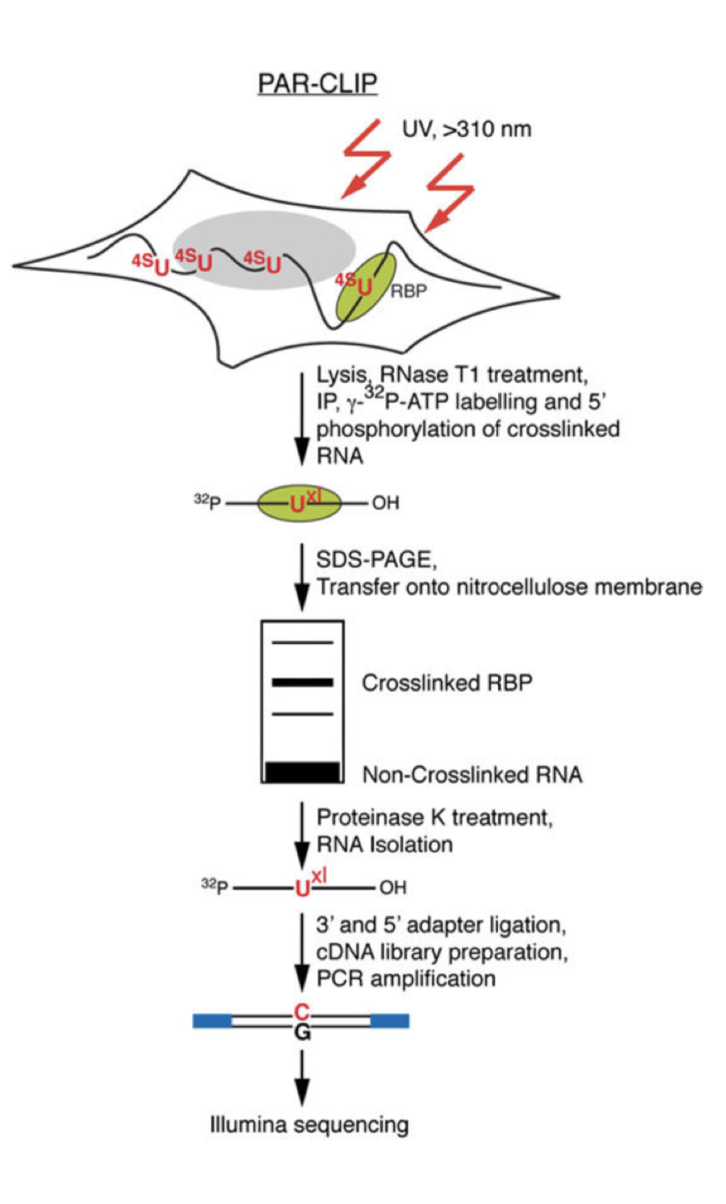
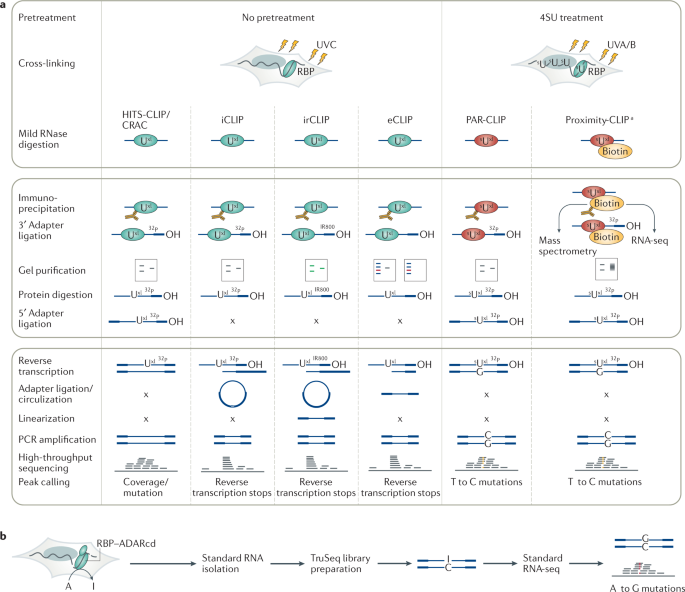
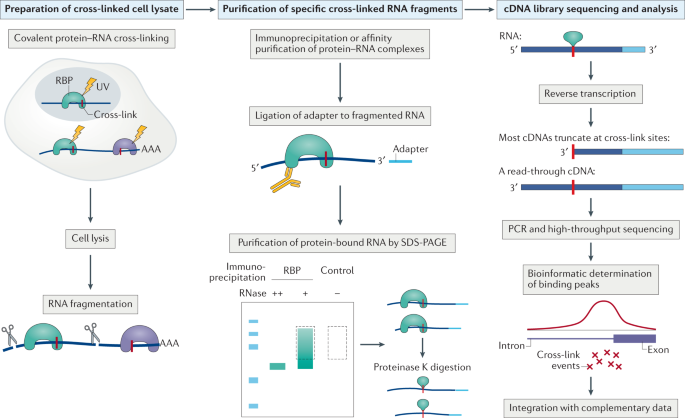
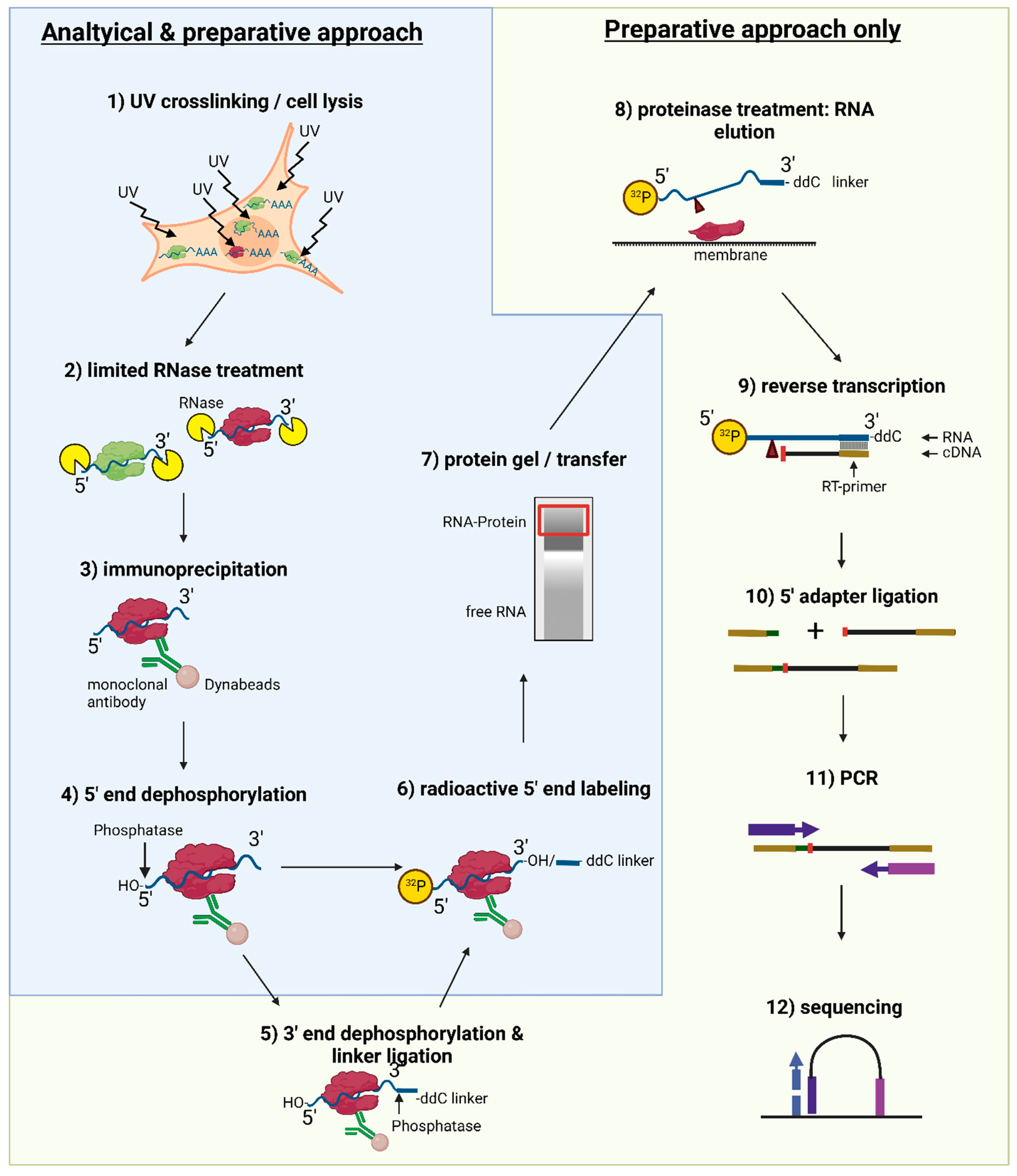
PAR-CLIP (Photoactivatable Ribonucleoside-Enhanced Crosslinking and Immunoprecipitation): a step-by-step protocol to the transcriptome-wide identification of binding sites of RNA-binding proteins. | Semantic Scholar
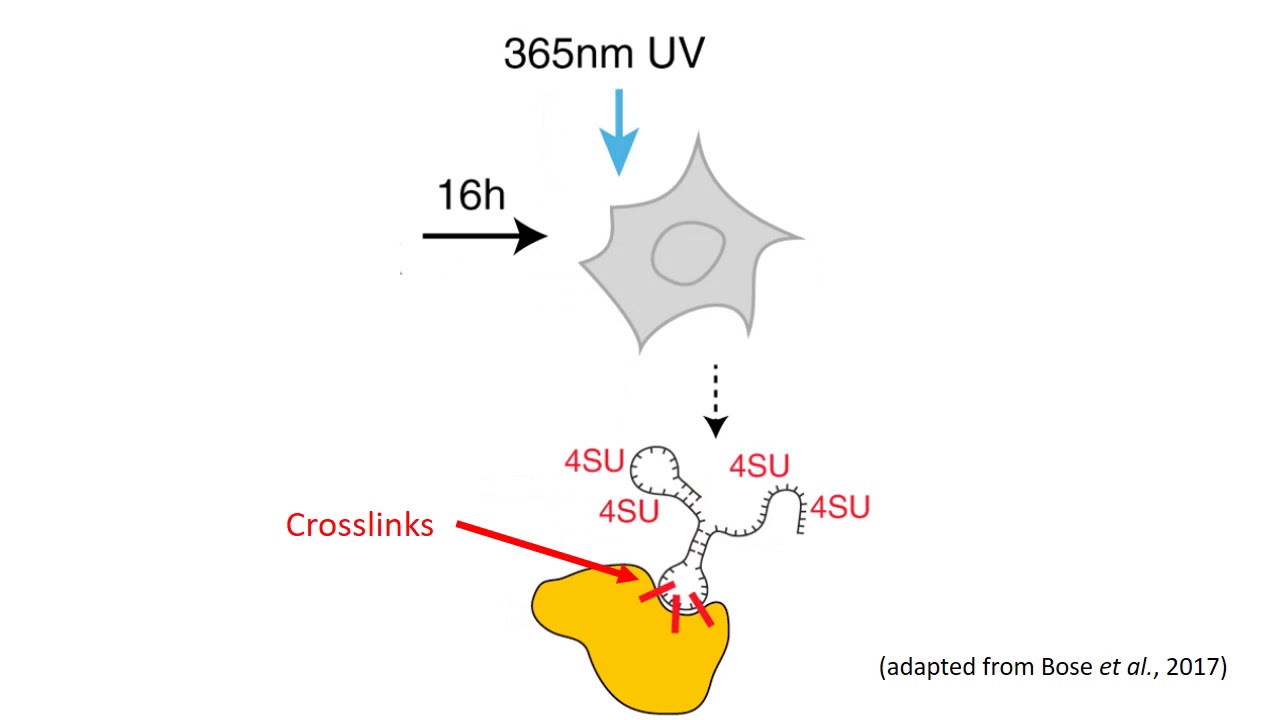
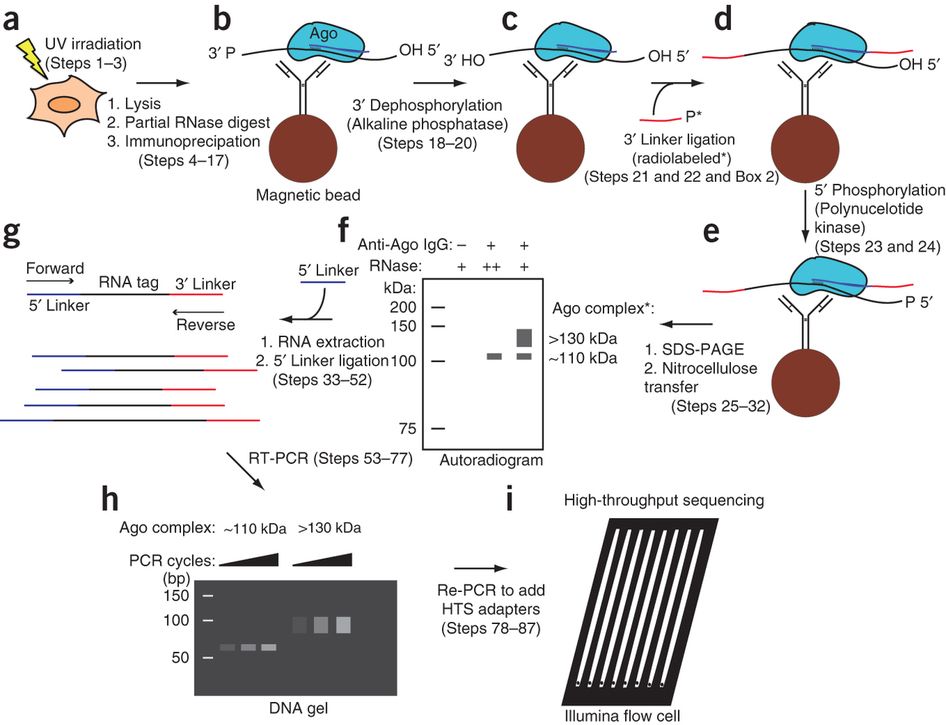
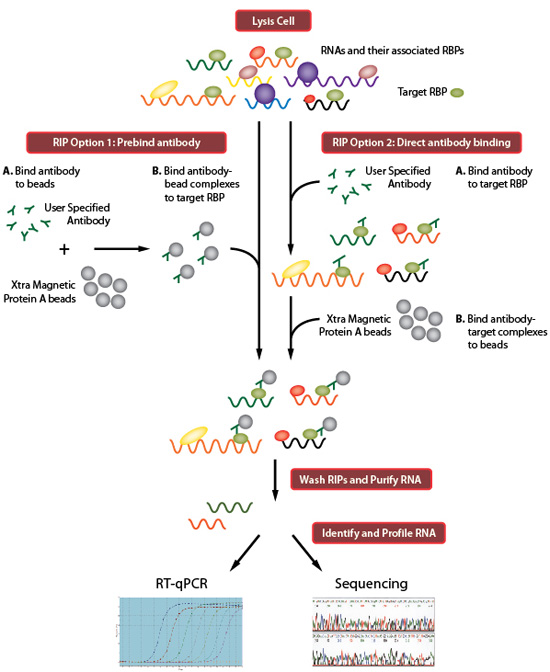
MPs | Free Full-Text | Studying miRNA–mRNA Interactions: An Optimized CLIP-Protocol for Endogenous Ago2-Protein

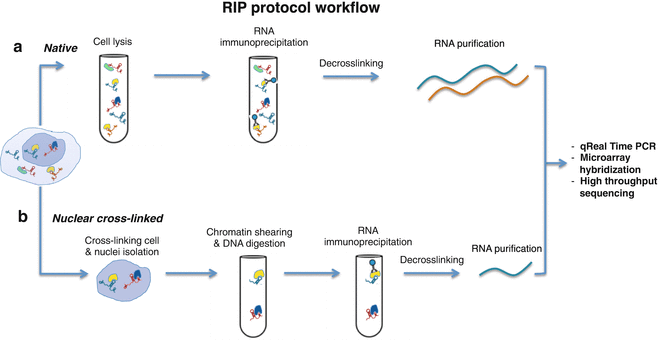
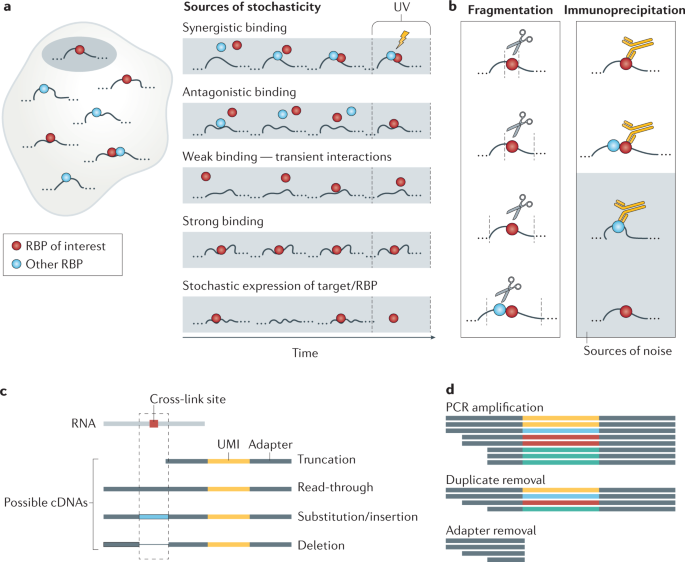
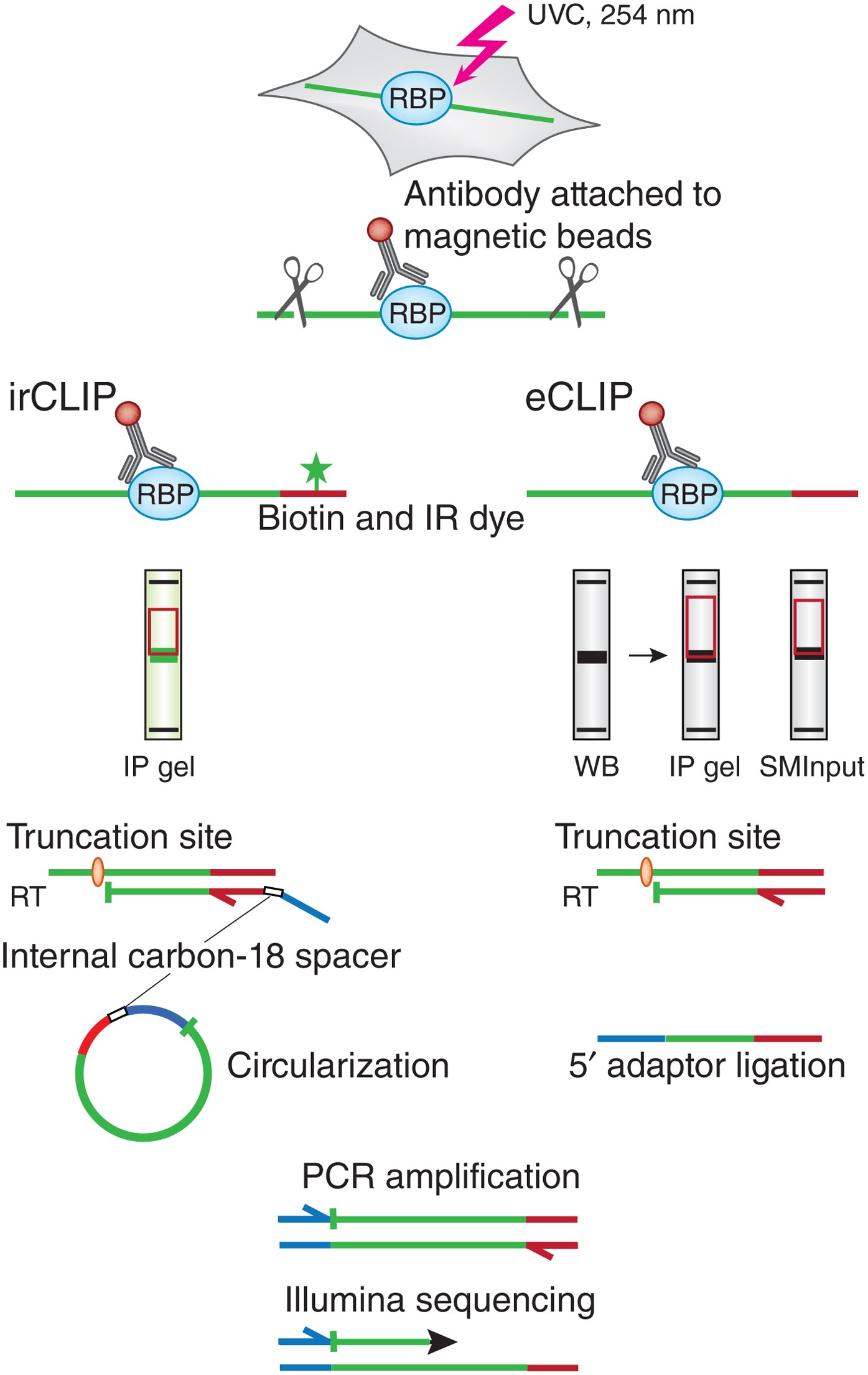
Computational approaches for the analysis of RNA–protein interactions: A primer for biologists - ScienceDirect

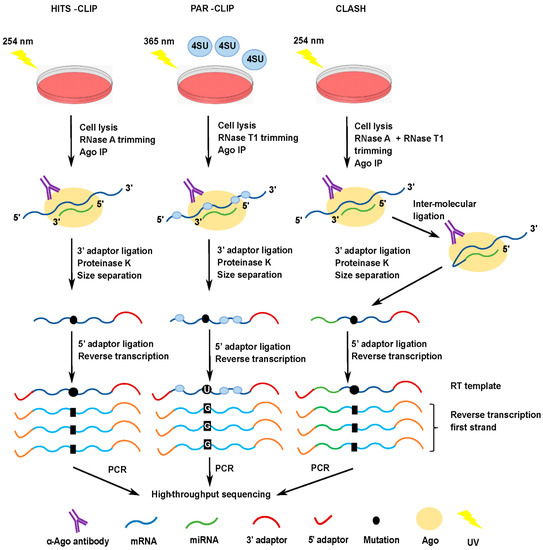
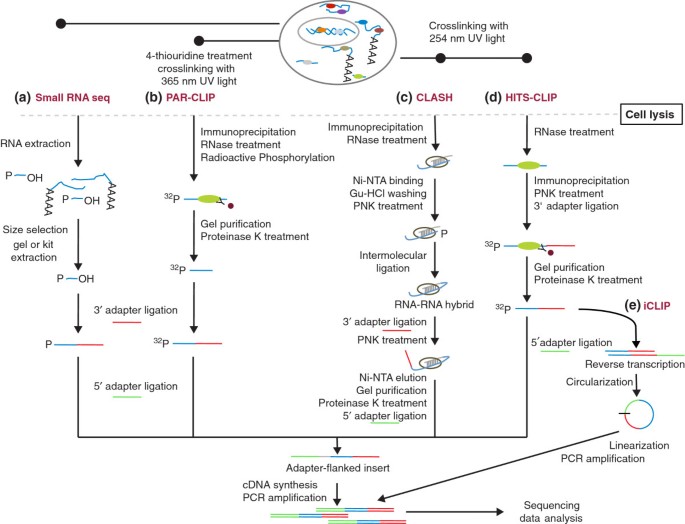
Outline of HITS-CLIP, PAR-CLIP and several variants, iCLIP, iCLAP and... | Download Scientific Diagram
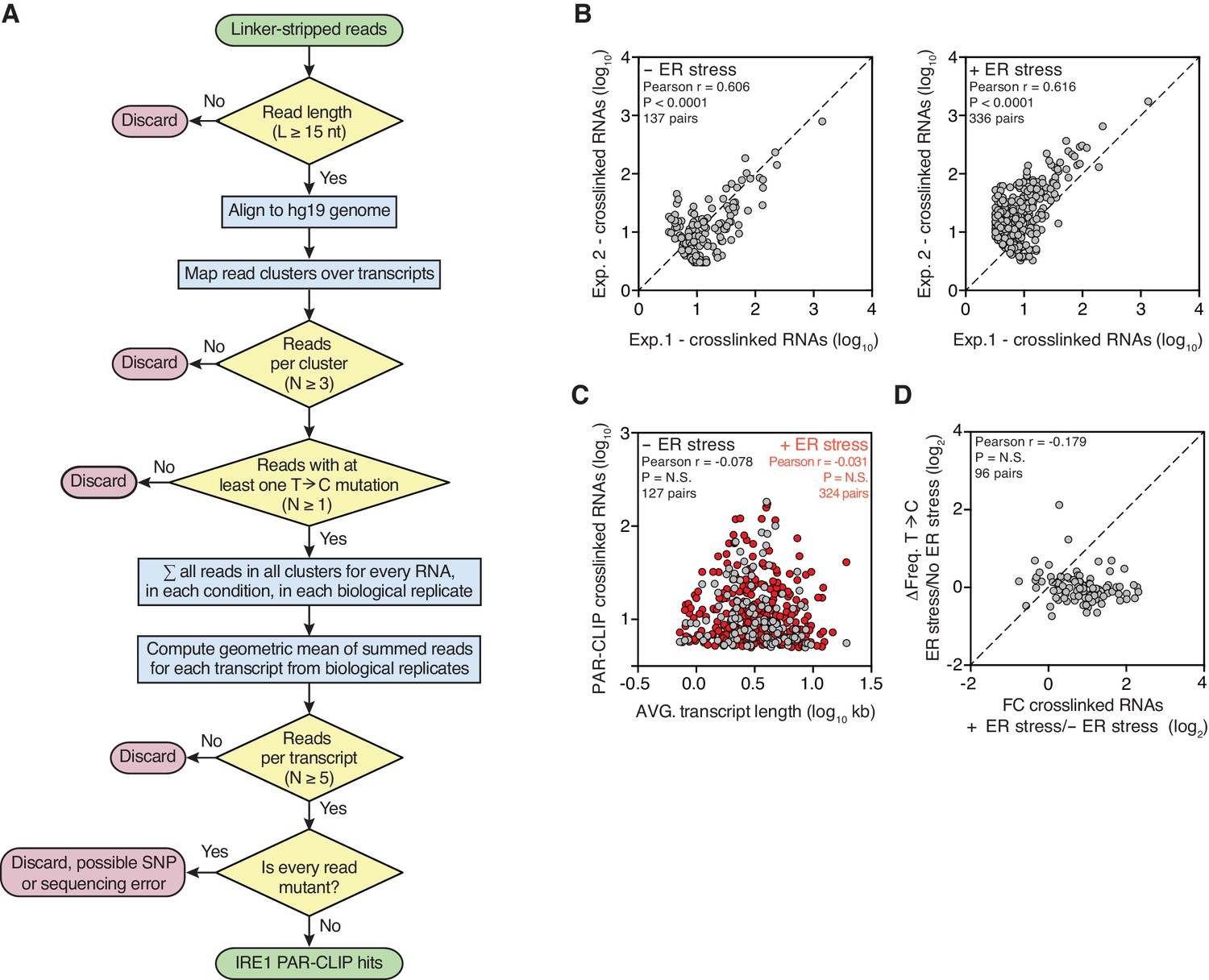
Figures and data in The unfolded protein response and endoplasmic reticulum protein targeting machineries converge on the stress sensor IRE1 | eLife

Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP: Cell

PAR-CLIP and Streamlined Small RNA cDNA Library Preparation Protocol for the Identification of RNA Binding Protein Target Sites | RNA-Seq Blog

Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP: Cell